Sodium »
PDB 3cq8-3dfh »
3ddk »
Sodium in PDB 3ddk: Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
Protein crystallography data
The structure of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase, PDB code: 3ddk
was solved by
G.Campagnola,
M.H.Weygandt,
K.E.Scoggin,
O.B.Peersen,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 52.58 / 2.25 |
| Space group | P 43 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 74.360, 74.360, 288.050, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 20.1 / 23.4 |
Sodium Binding Sites:
The binding sites of Sodium atom in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
(pdb code 3ddk). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total 9 binding sites of Sodium where determined in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase, PDB code: 3ddk:
Jump to Sodium binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9;
In total 9 binding sites of Sodium where determined in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase, PDB code: 3ddk:
Jump to Sodium binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9;
Sodium binding site 1 out of 9 in 3ddk
Go back to
Sodium binding site 1 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
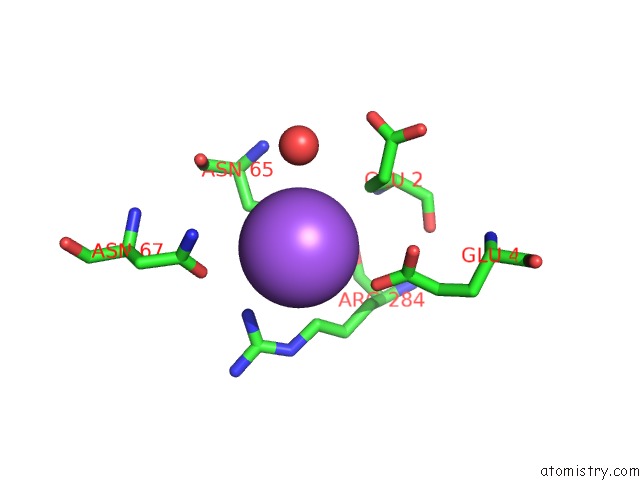
Mono view
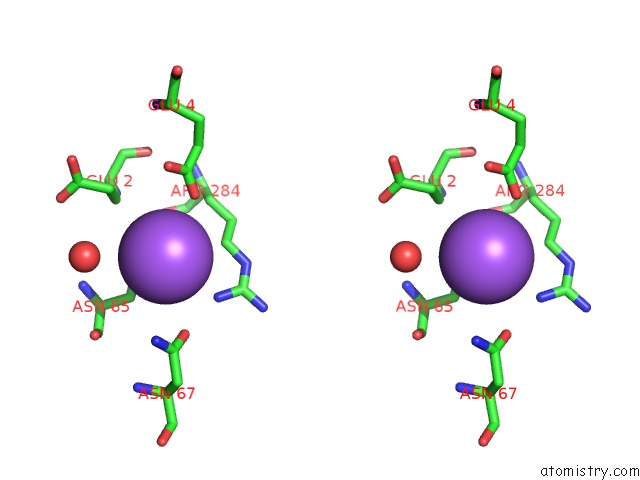
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 2 out of 9 in 3ddk
Go back to
Sodium binding site 2 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
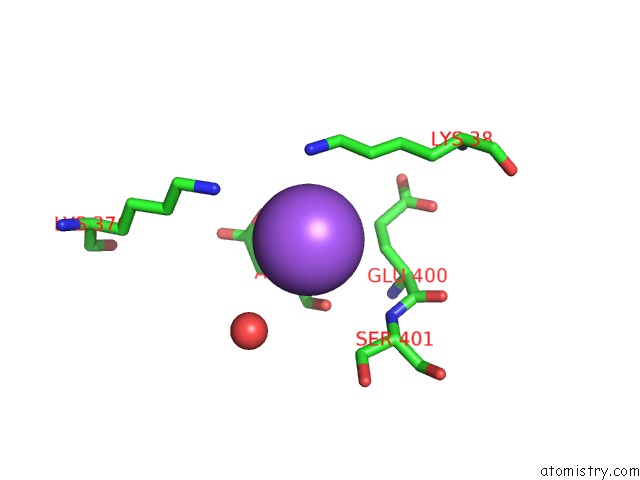
Mono view
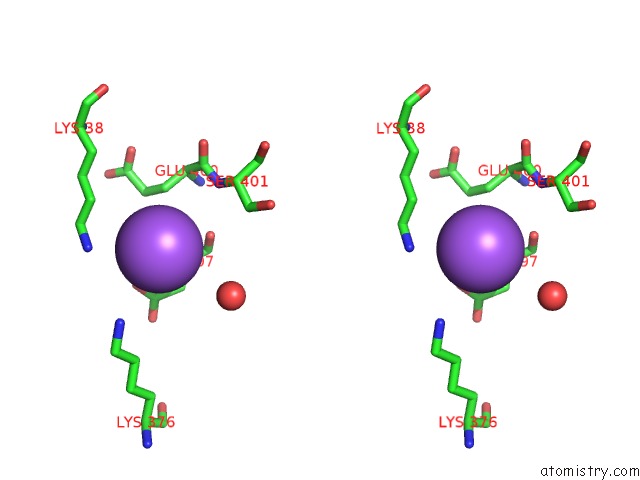
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 2 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 3 out of 9 in 3ddk
Go back to
Sodium binding site 3 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
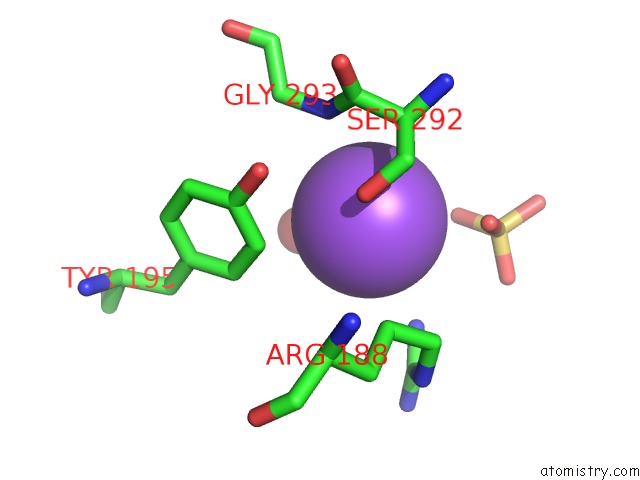
Mono view
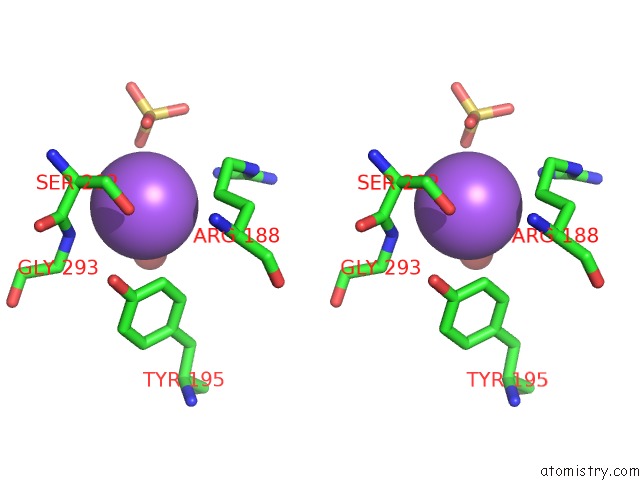
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 3 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 4 out of 9 in 3ddk
Go back to
Sodium binding site 4 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
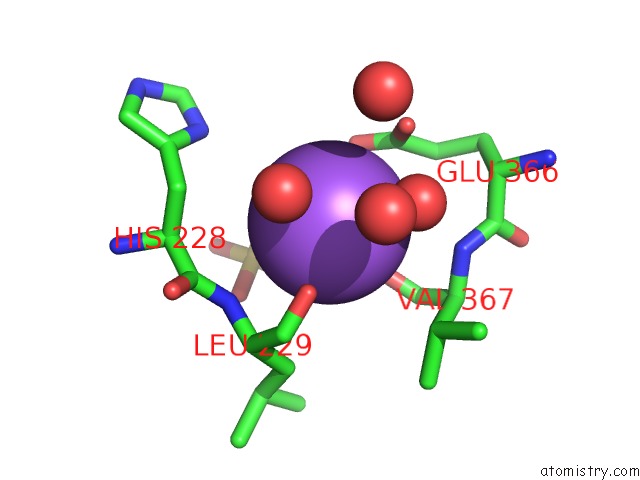
Mono view
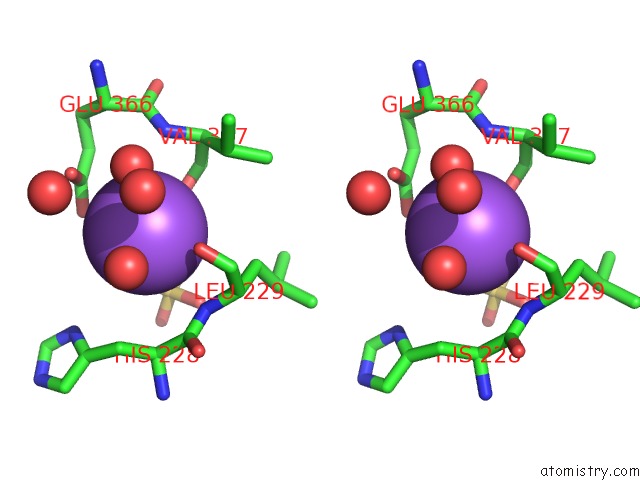
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 4 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 5 out of 9 in 3ddk
Go back to
Sodium binding site 5 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
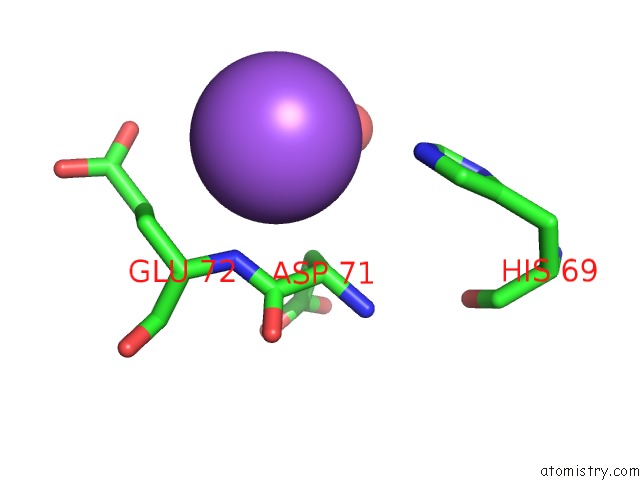
Mono view
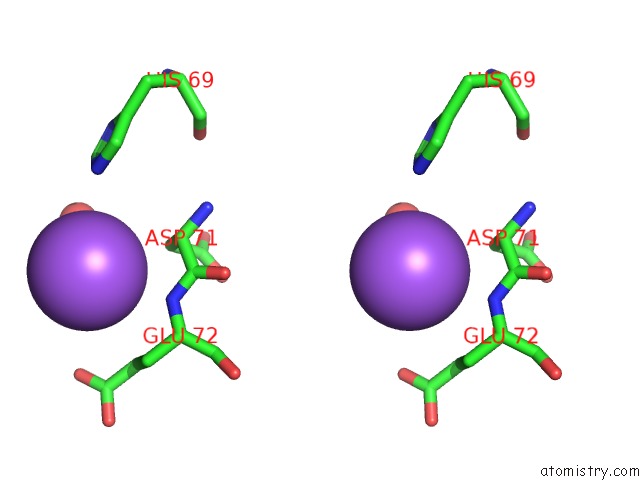
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 5 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 6 out of 9 in 3ddk
Go back to
Sodium binding site 6 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
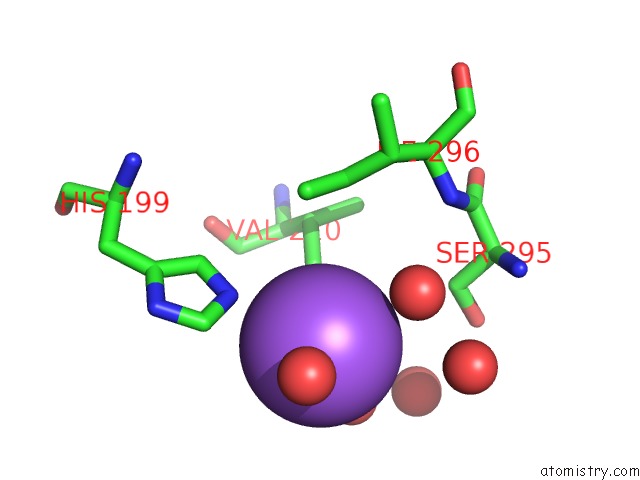
Mono view
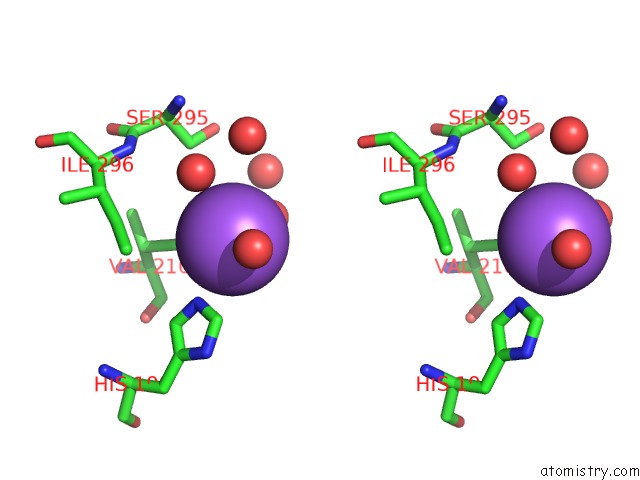
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 6 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 7 out of 9 in 3ddk
Go back to
Sodium binding site 7 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
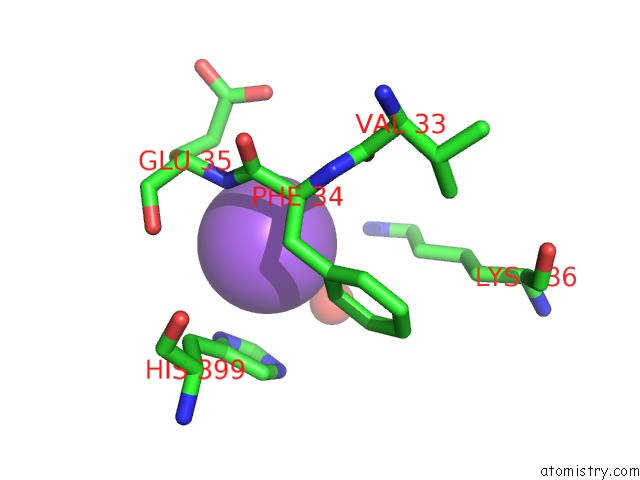
Mono view
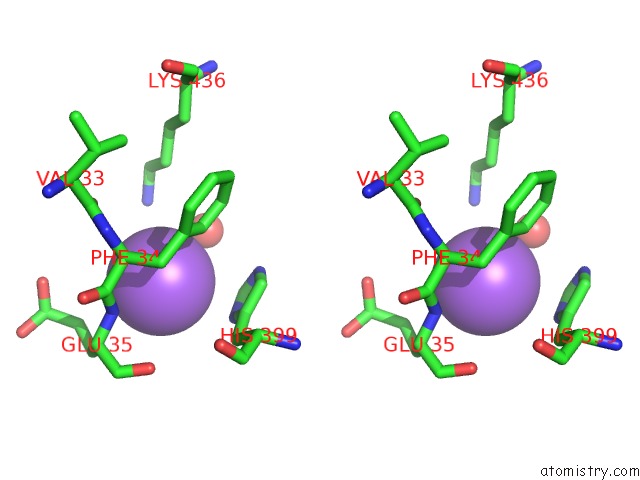
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 7 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 8 out of 9 in 3ddk
Go back to
Sodium binding site 8 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
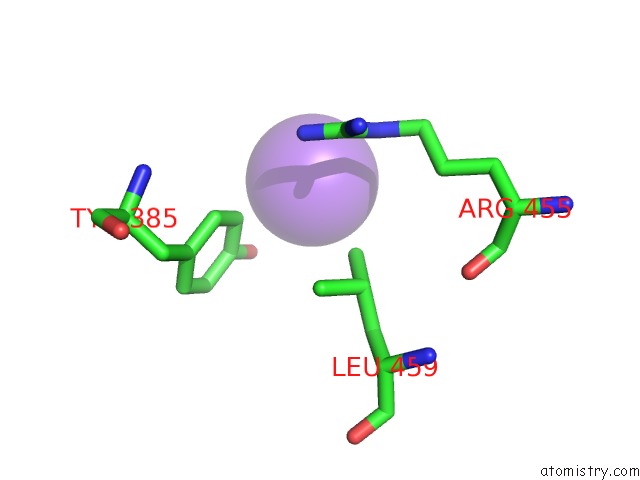
Mono view
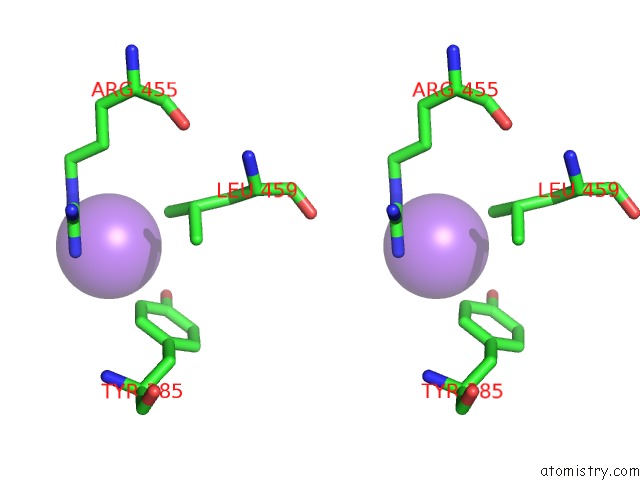
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 8 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Sodium binding site 9 out of 9 in 3ddk
Go back to
Sodium binding site 9 out
of 9 in the Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase
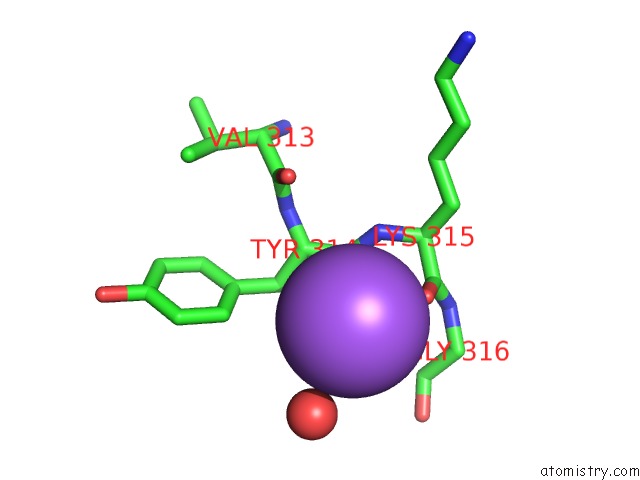
Mono view
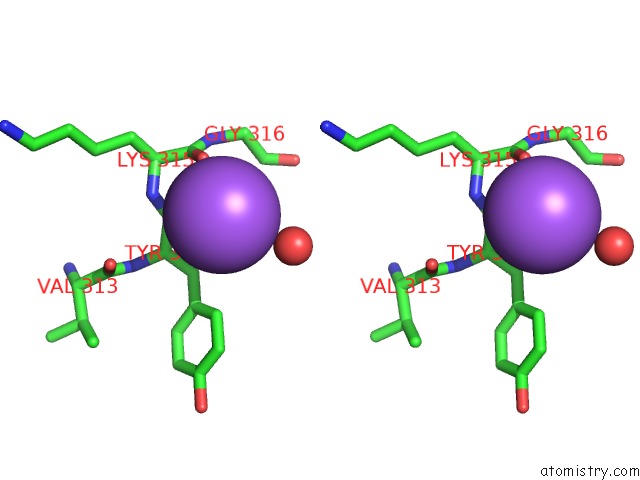
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 9 of Coxsackievirus B3 3DPOL Rna Dependent Rna Polymerase within 5.0Å range:
|
Reference:
G.Campagnola,
M.H.Weygandt,
K.E.Scoggin,
O.B.Peersen.
Crystal Structure of Coxsackievirus B3 3DPOL Highlights Functional Importance of Residue 5 in Picornaviral Polymerases J.Virol. V. 82 9458 2008.
ISSN: ISSN 0022-538X
PubMed: 18632862
DOI: 10.1128/JVI.00647-08
Page generated: Sun Aug 17 14:20:05 2025
ISSN: ISSN 0022-538X
PubMed: 18632862
DOI: 10.1128/JVI.00647-08
Last articles
Na in 6J6VNa in 6J6O
Na in 6J3B
Na in 6J05
Na in 6J0H
Na in 6IWH
Na in 6IX8
Na in 6IX9
Na in 6IWG
Na in 6IX5