Sodium »
PDB 5ack-5b07 »
5ahg »
Sodium in PDB 5ahg: Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine
Enzymatic activity of Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine
All present enzymatic activity of Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine:
3.4.21.5;
3.4.21.5;
Protein crystallography data
The structure of Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine, PDB code: 5ahg
was solved by
E.Ruehmann,
A.Heine,
G.Klebe,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 43.41 / 1.24 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 70.222, 71.263, 72.216, 90.00, 100.48, 90.00 |
| R / Rfree (%) | 12.3 / 14.4 |
Other elements in 5ahg:
The structure of Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine also contains other interesting chemical elements:
| Chlorine | (Cl) | 1 atom |
Sodium Binding Sites:
The binding sites of Sodium atom in the Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine
(pdb code 5ahg). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total 2 binding sites of Sodium where determined in the Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine, PDB code: 5ahg:
Jump to Sodium binding site number: 1; 2;
In total 2 binding sites of Sodium where determined in the Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine, PDB code: 5ahg:
Jump to Sodium binding site number: 1; 2;
Sodium binding site 1 out of 2 in 5ahg
Go back to
Sodium binding site 1 out
of 2 in the Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine
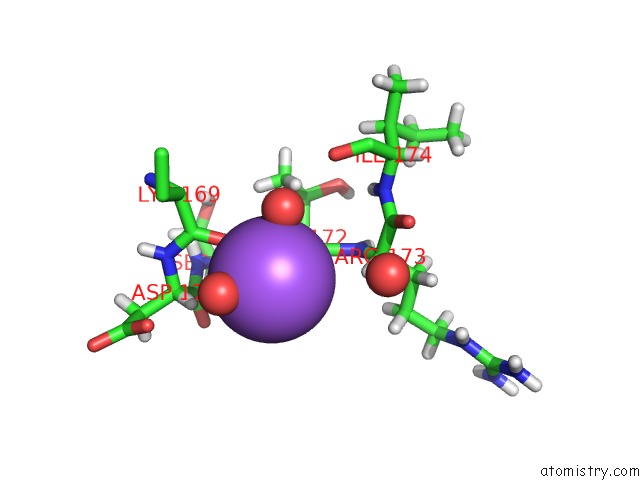
Mono view
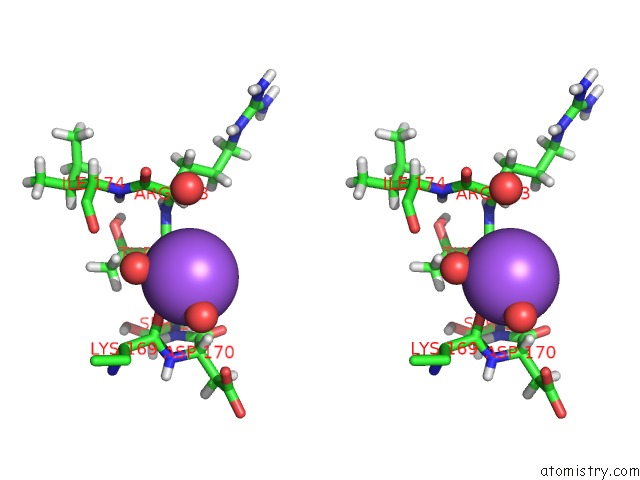
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine within 5.0Å range:
|
Sodium binding site 2 out of 2 in 5ahg
Go back to
Sodium binding site 2 out
of 2 in the Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine
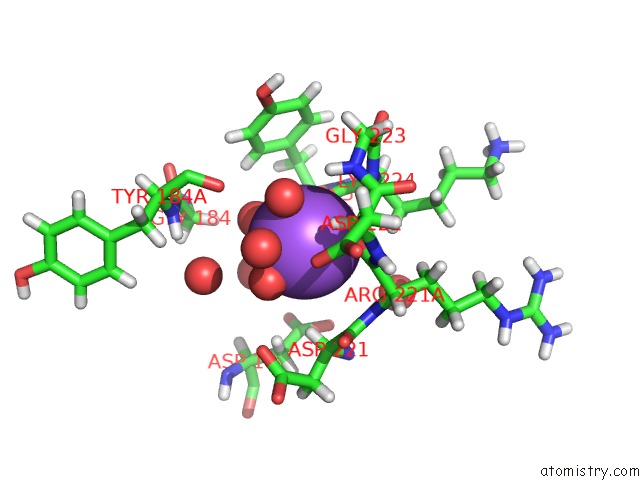
Mono view
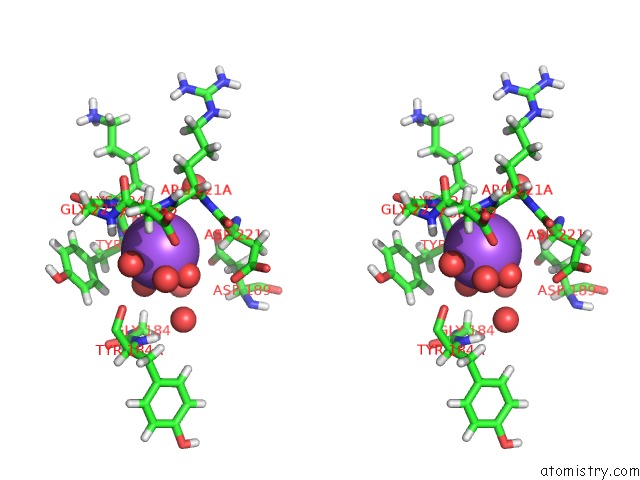
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 2 of Thrombin in Complex with ((4-Chlorophenyl)Sulfamoyl))Diemethylamine within 5.0Å range:
|
Reference:
E.Ruehmann,
M.Betz,
A.Heine,
G.Klebe.
Fragments Can Bind Either More Enthalpy or Entropy-Driven: Crystal Structures and Residual Hydration Pattern Suggest Why. J.Med.Chem. V. 58 6960 2015.
ISSN: ISSN 0022-2623
PubMed: 26270568
DOI: 10.1021/ACS.JMEDCHEM.5B00812
Page generated: Mon Oct 7 19:54:12 2024
ISSN: ISSN 0022-2623
PubMed: 26270568
DOI: 10.1021/ACS.JMEDCHEM.5B00812
Last articles
Cl in 5SE3Cl in 5SDS
Cl in 5SEE
Cl in 5SDR
Cl in 5SDQ
Cl in 5SDP
Cl in 5SDN
Cl in 5SDO
Cl in 5SDM
Cl in 5SDL