Sodium »
PDB 1jb7-1k73 »
1jlj »
Sodium in PDB 1jlj: 1.6 Angstrom Crystal Structure of the Human Neuroreceptor Anchoring and Molybdenum Cofactor Biosynthesis Protein Gephyrin
Protein crystallography data
The structure of 1.6 Angstrom Crystal Structure of the Human Neuroreceptor Anchoring and Molybdenum Cofactor Biosynthesis Protein Gephyrin, PDB code: 1jlj
was solved by
G.Schwarz,
N.Schrader,
R.R.Mendel,
H.-J.Hecht,
H.Schindelin,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 20.00 / 1.60 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 114.269, 66.008, 73.122, 90.00, 93.27, 90.00 |
| R / Rfree (%) | 14.7 / 18.3 |
Sodium Binding Sites:
The binding sites of Sodium atom in the 1.6 Angstrom Crystal Structure of the Human Neuroreceptor Anchoring and Molybdenum Cofactor Biosynthesis Protein Gephyrin
(pdb code 1jlj). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total only one binding site of Sodium was determined in the 1.6 Angstrom Crystal Structure of the Human Neuroreceptor Anchoring and Molybdenum Cofactor Biosynthesis Protein Gephyrin, PDB code: 1jlj:
In total only one binding site of Sodium was determined in the 1.6 Angstrom Crystal Structure of the Human Neuroreceptor Anchoring and Molybdenum Cofactor Biosynthesis Protein Gephyrin, PDB code: 1jlj:
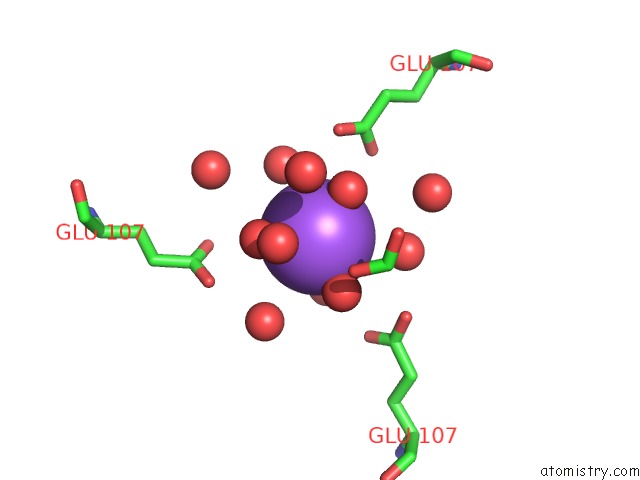
Sodium binding site 1 out of 1 in 1jlj
Go back to
Sodium binding site 1 out
of 1 in the 1.6 Angstrom Crystal Structure of the Human Neuroreceptor Anchoring and Molybdenum Cofactor Biosynthesis Protein Gephyrin
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of 1.6 Angstrom Crystal Structure of the Human Neuroreceptor Anchoring and Molybdenum Cofactor Biosynthesis Protein Gephyrin within 5.0Å range:
|
Reference:
G.Schwarz,
N.Schrader,
R.R.Mendel,
H.J.Hecht,
H.Schindelin.
Crystal Structures of Human Gephyrin and Plant CNX1 G Domains: Comparative Analysis and Functional Implications. J.Mol.Biol. V. 312 405 2001.
ISSN: ISSN 0022-2836
PubMed: 11554796
DOI: 10.1006/JMBI.2001.4952
Page generated: Sun Aug 17 05:20:56 2025
ISSN: ISSN 0022-2836
PubMed: 11554796
DOI: 10.1006/JMBI.2001.4952
Last articles
Na in 5INBNa in 5IMF
Na in 5IMZ
Na in 5IMV
Na in 5IM9
Na in 5IMD
Na in 5IMC
Na in 5IMA
Na in 5IIZ
Na in 5IIO