Sodium »
PDB 4owe-4pd7 »
4p30 »
Sodium in PDB 4p30: Structure of Navms Mutant in Presence of PI1 Compound
Protein crystallography data
The structure of Structure of Navms Mutant in Presence of PI1 Compound, PDB code: 4p30
was solved by
C.Bagneris,
C.E.Naylor,
B.A.Wallace,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 45.73 / 3.31 |
| Space group | C 2 2 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 80.260, 334.260, 80.040, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 20.4 / 22.5 |
Sodium Binding Sites:
The binding sites of Sodium atom in the Structure of Navms Mutant in Presence of PI1 Compound
(pdb code 4p30). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total 3 binding sites of Sodium where determined in the Structure of Navms Mutant in Presence of PI1 Compound, PDB code: 4p30:
Jump to Sodium binding site number: 1; 2; 3;
In total 3 binding sites of Sodium where determined in the Structure of Navms Mutant in Presence of PI1 Compound, PDB code: 4p30:
Jump to Sodium binding site number: 1; 2; 3;
Sodium binding site 1 out of 3 in 4p30
Go back to
Sodium binding site 1 out
of 3 in the Structure of Navms Mutant in Presence of PI1 Compound
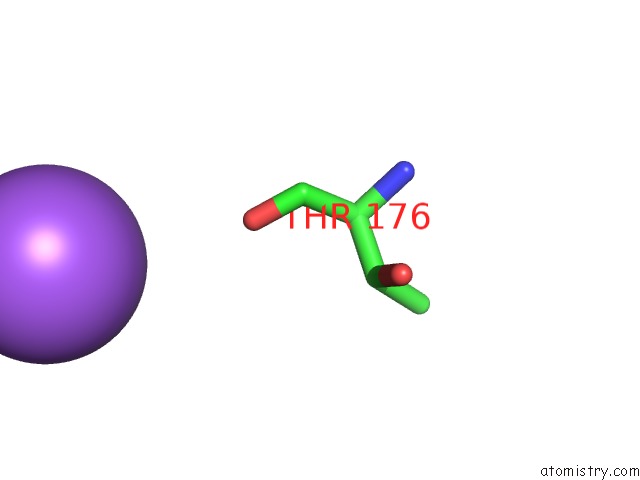
Mono view
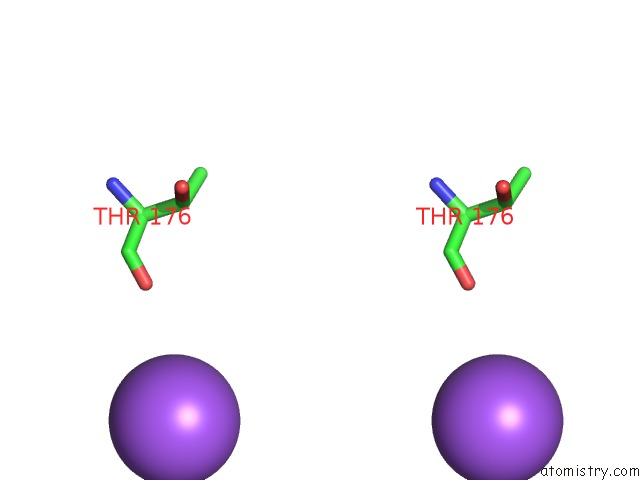
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of Structure of Navms Mutant in Presence of PI1 Compound within 5.0Å range:
|
Sodium binding site 2 out of 3 in 4p30
Go back to
Sodium binding site 2 out
of 3 in the Structure of Navms Mutant in Presence of PI1 Compound
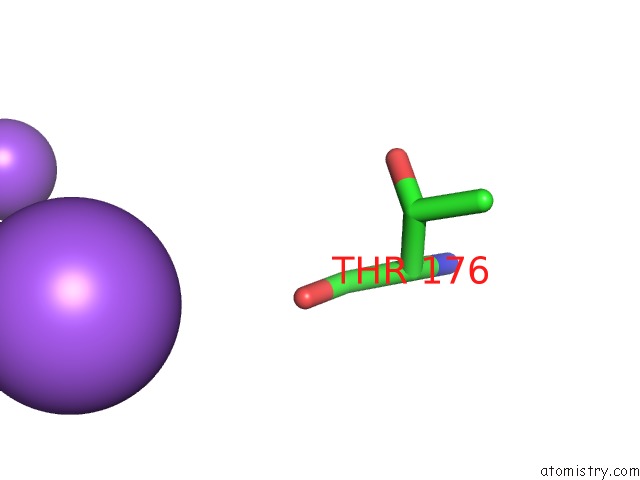
Mono view
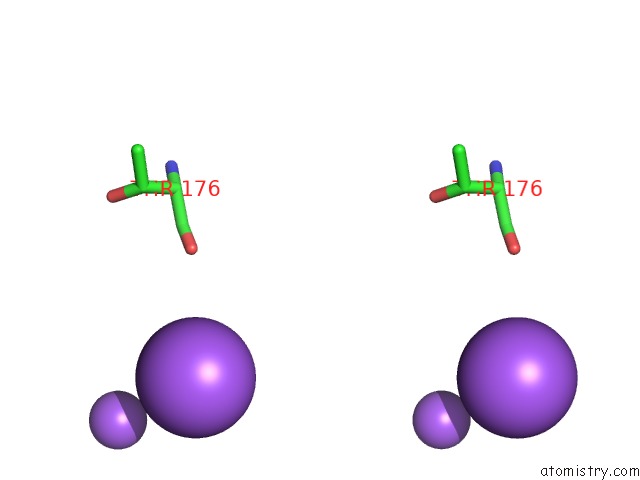
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 2 of Structure of Navms Mutant in Presence of PI1 Compound within 5.0Å range:
|
Sodium binding site 3 out of 3 in 4p30
Go back to
Sodium binding site 3 out
of 3 in the Structure of Navms Mutant in Presence of PI1 Compound
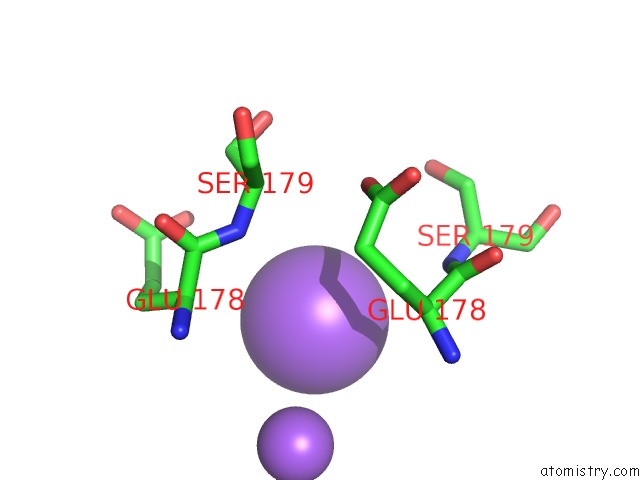
Mono view
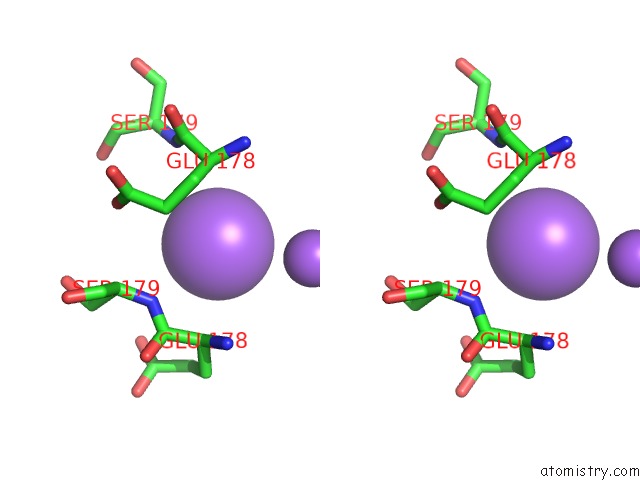
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 3 of Structure of Navms Mutant in Presence of PI1 Compound within 5.0Å range:
|
Reference:
C.Bagneris,
P.G.Decaen,
C.E.Naylor,
D.C.Pryde,
I.Nobeli,
D.E.Clapham,
B.A.Wallace.
Prokaryotic Navms Channel As A Structural and Functional Model For Eukaryotic Sodium Channel Antagonism. Proc.Natl.Acad.Sci.Usa V. 111 8428 2014.
ISSN: ESSN 1091-6490
PubMed: 24850863
DOI: 10.1073/PNAS.1406855111
Page generated: Mon Oct 7 17:36:54 2024
ISSN: ESSN 1091-6490
PubMed: 24850863
DOI: 10.1073/PNAS.1406855111
Last articles
Mg in 1S8FMg in 1S83
Mg in 1S77
Mg in 1S76
Mg in 1S4E
Mg in 1S6P
Mg in 1S6H
Mg in 1S5J
Mg in 1S5G
Mg in 1S2V