Sodium »
PDB 5ppq-5pqt »
5pq4 »
Sodium in PDB 5pq4: Pandda Analysis Group Deposition -- Crystal Structure of BRD1 After Initial Refinement with No Ligand Modelled (Structure 41)
Protein crystallography data
The structure of Pandda Analysis Group Deposition -- Crystal Structure of BRD1 After Initial Refinement with No Ligand Modelled (Structure 41), PDB code: 5pq4
was solved by
N.M.Pearce,
T.Krojer,
R.Talon,
A.R.Bradley,
M.Fairhead,
R.Sethi,
N.Wright,
E.Maclean,
P.Collins,
J.Brandao-Neto,
A.Douangamath,
Z.Renjie,
A.Dias,
J.Ng,
P.E.Brennan,
O.Cox,
C.Bountra,
C.H.Arrowsmith,
A.Edwards,
F.Vondelft,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 50.86 / 1.63 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 55.090, 56.350, 101.610, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 18.4 / 22.4 |
Sodium Binding Sites:
The binding sites of Sodium atom in the Pandda Analysis Group Deposition -- Crystal Structure of BRD1 After Initial Refinement with No Ligand Modelled (Structure 41)
(pdb code 5pq4). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total only one binding site of Sodium was determined in the Pandda Analysis Group Deposition -- Crystal Structure of BRD1 After Initial Refinement with No Ligand Modelled (Structure 41), PDB code: 5pq4:
In total only one binding site of Sodium was determined in the Pandda Analysis Group Deposition -- Crystal Structure of BRD1 After Initial Refinement with No Ligand Modelled (Structure 41), PDB code: 5pq4:
Sodium binding site 1 out of 1 in 5pq4
Go back to
Sodium binding site 1 out
of 1 in the Pandda Analysis Group Deposition -- Crystal Structure of BRD1 After Initial Refinement with No Ligand Modelled (Structure 41)
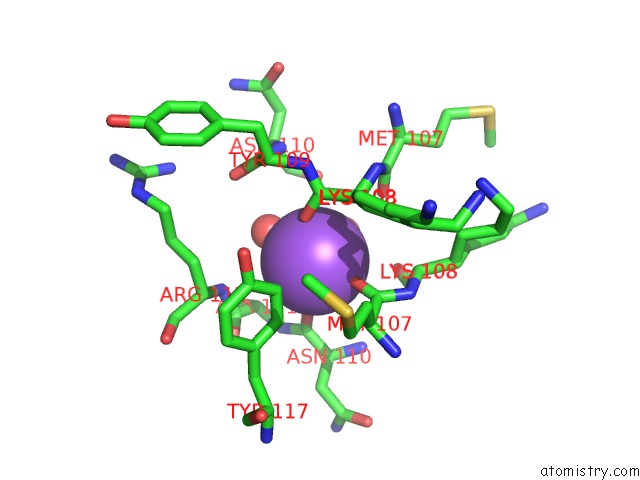
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of Pandda Analysis Group Deposition -- Crystal Structure of BRD1 After Initial Refinement with No Ligand Modelled (Structure 41) within 5.0Å range:
|
Reference:
N.M.Pearce,
T.Krojer,
A.R.Bradley,
P.Collins,
R.P.Nowak,
R.Talon,
B.D.Marsden,
S.Kelm,
J.Shi,
C.M.Deane,
F.Von Delft.
A Multi-Crystal Method For Extracting Obscured Crystallographic States From Conventionally Uninterpretable Electron Density. Nat Commun V. 8 15123 2017.
ISSN: ESSN 2041-1723
PubMed: 28436492
DOI: 10.1038/NCOMMS15123
Page generated: Mon Oct 7 23:27:02 2024
ISSN: ESSN 2041-1723
PubMed: 28436492
DOI: 10.1038/NCOMMS15123
Last articles
Cl in 7WGOCl in 7WGD
Cl in 7WFR
Cl in 7WF8
Cl in 7WEL
Cl in 7WE4
Cl in 7WCX
Cl in 7WCW
Cl in 7WCT
Cl in 7WCV