Sodium »
PDB 5mss-5nbj »
5mzq »
Sodium in PDB 5mzq: X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
Protein crystallography data
The structure of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform, PDB code: 5mzq
was solved by
Z.Fourati,
M.Delarue,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 19.84 / 2.80 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 181.969, 131.779, 160.404, 90.00, 102.93, 90.00 |
| R / Rfree (%) | 20.2 / 21.9 |
Other elements in 5mzq:
The structure of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform also contains other interesting chemical elements:
| Bromine | (Br) | 15 atoms |
| Chlorine | (Cl) | 7 atoms |
Sodium Binding Sites:
The binding sites of Sodium atom in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
(pdb code 5mzq). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total 6 binding sites of Sodium where determined in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform, PDB code: 5mzq:
Jump to Sodium binding site number: 1; 2; 3; 4; 5; 6;
In total 6 binding sites of Sodium where determined in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform, PDB code: 5mzq:
Jump to Sodium binding site number: 1; 2; 3; 4; 5; 6;
Sodium binding site 1 out of 6 in 5mzq
Go back to
Sodium binding site 1 out
of 6 in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
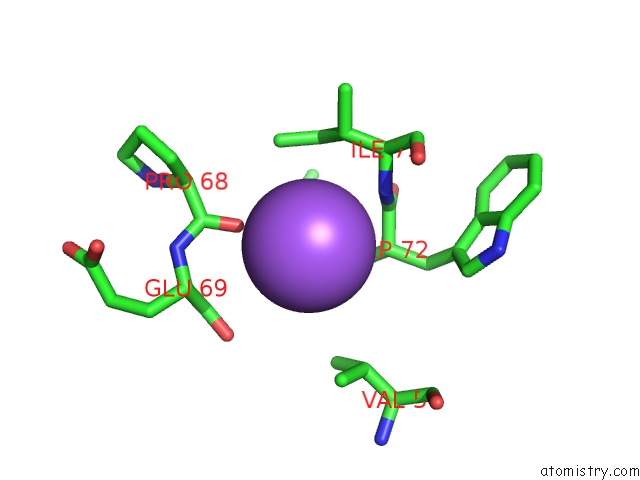
Mono view
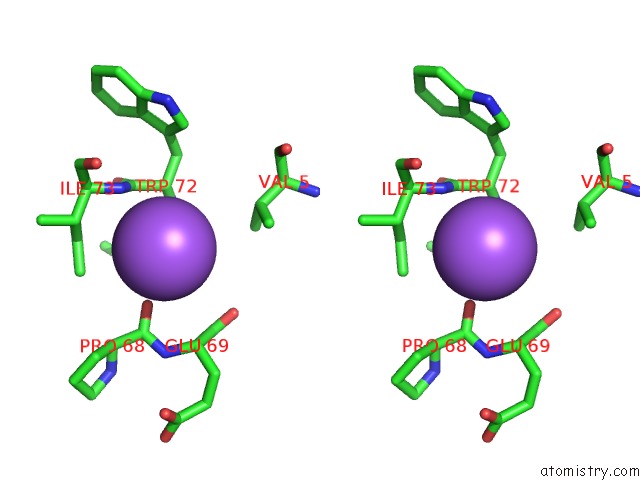
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform within 5.0Å range:
|
Sodium binding site 2 out of 6 in 5mzq
Go back to
Sodium binding site 2 out
of 6 in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
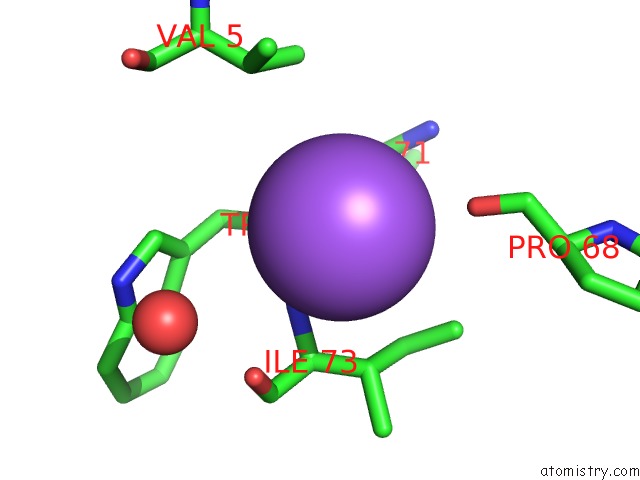
Mono view
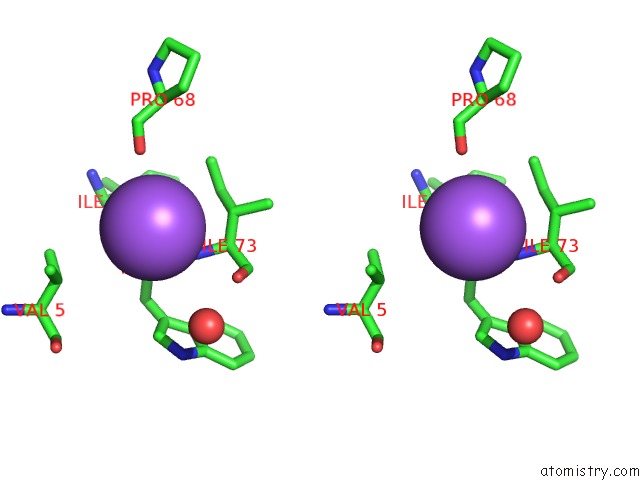
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 2 of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform within 5.0Å range:
|
Sodium binding site 3 out of 6 in 5mzq
Go back to
Sodium binding site 3 out
of 6 in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
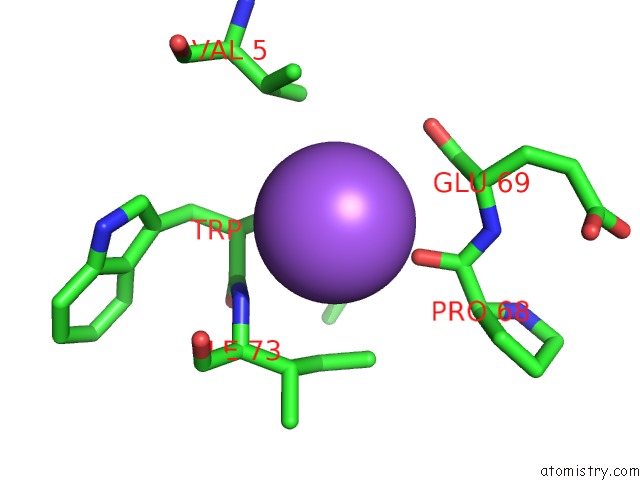
Mono view
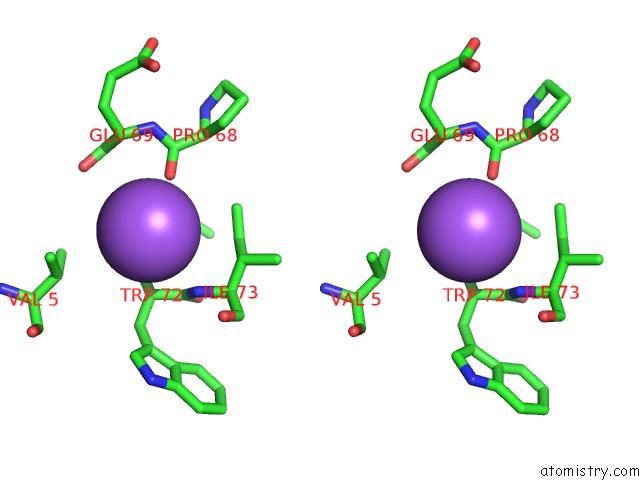
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 3 of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform within 5.0Å range:
|
Sodium binding site 4 out of 6 in 5mzq
Go back to
Sodium binding site 4 out
of 6 in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
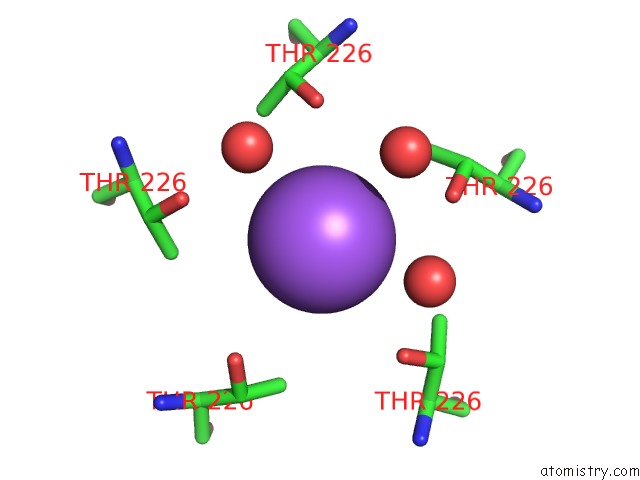
Mono view
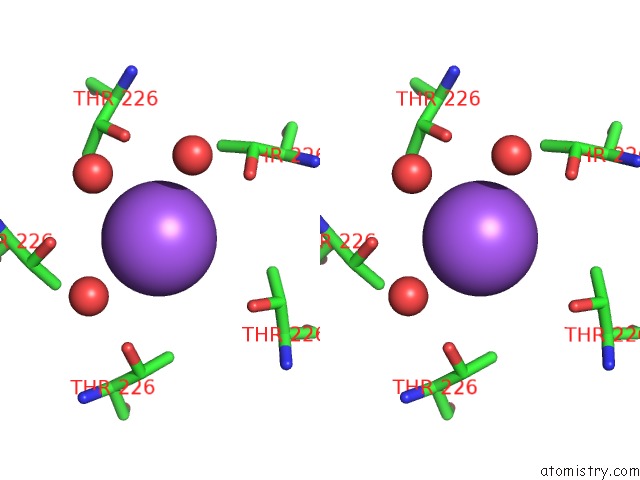
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 4 of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform within 5.0Å range:
|
Sodium binding site 5 out of 6 in 5mzq
Go back to
Sodium binding site 5 out
of 6 in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
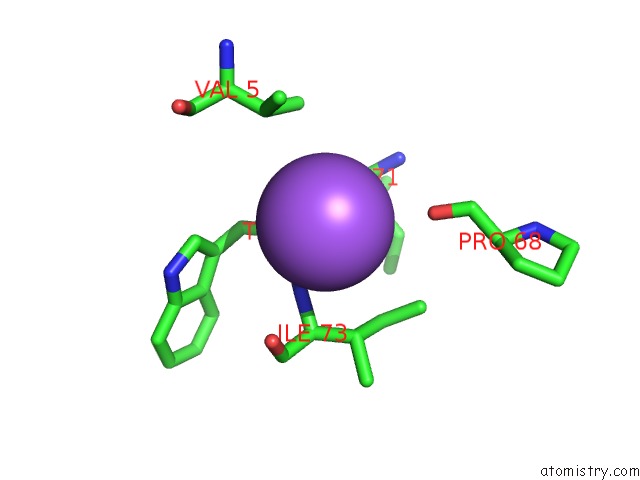
Mono view
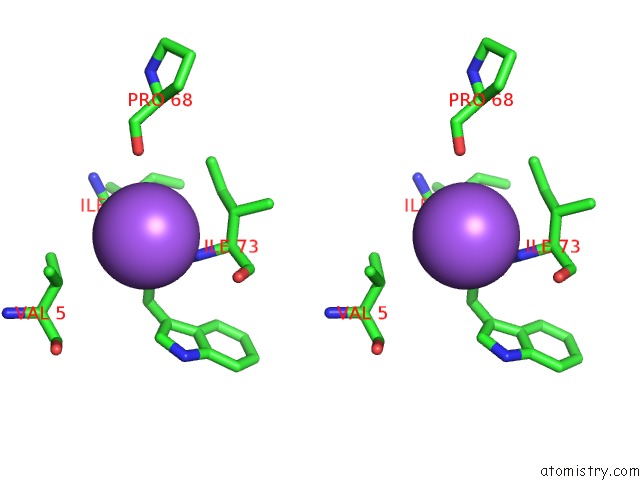
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 5 of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform within 5.0Å range:
|
Sodium binding site 6 out of 6 in 5mzq
Go back to
Sodium binding site 6 out
of 6 in the X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform
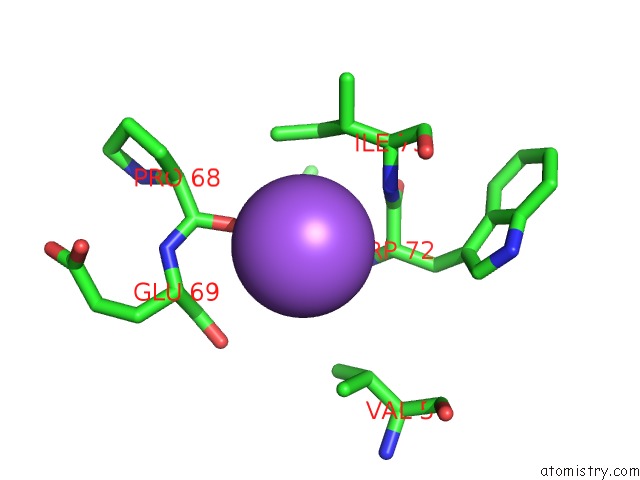
Mono view
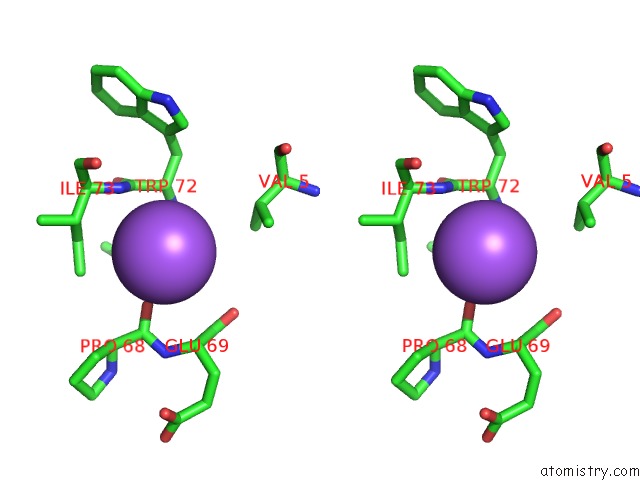
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 6 of X-Ray Structure of the M205W Mutant of Glic in Complex with Bromoform within 5.0Å range:
|
Reference:
Z.Fourati,
R.J.Howard,
S.A.Heusser,
H.Hu,
R.R.Ruza,
L.Sauguet,
E.Lindahl,
M.Delarue.
Structural Basis For A Bimodal Allosteric Mechanism of General Anesthetic Modulation in Pentameric Ligand-Gated Ion Channels. Cell Rep V. 23 993 2018.
ISSN: ESSN 2211-1247
PubMed: 29694907
DOI: 10.1016/J.CELREP.2018.03.108
Page generated: Mon Oct 7 22:49:37 2024
ISSN: ESSN 2211-1247
PubMed: 29694907
DOI: 10.1016/J.CELREP.2018.03.108
Last articles
Cl in 7VSLCl in 7VSI
Cl in 7VQM
Cl in 7VR9
Cl in 7VPE
Cl in 7VRE
Cl in 7VRA
Cl in 7VP8
Cl in 7VOE
Cl in 7VOD