Sodium »
PDB 3q11-3qpz »
3qng »
Sodium in PDB 3qng: Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)]
Enzymatic activity of Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)]
All present enzymatic activity of Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)]:
3.2.1.17;
3.2.1.17;
Protein crystallography data
The structure of Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)], PDB code: 3qng
was solved by
F.Zobi,
B.Spingler,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 33.12 / 1.55 |
| Space group | P 43 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 78.343, 78.343, 36.546, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 18.5 / 21.7 |
Other elements in 3qng:
The structure of Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)] also contains other interesting chemical elements:
| Rhenium | (Re) | 1 atom |
| Chlorine | (Cl) | 1 atom |
Sodium Binding Sites:
The binding sites of Sodium atom in the Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)]
(pdb code 3qng). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total only one binding site of Sodium was determined in the Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)], PDB code: 3qng:
In total only one binding site of Sodium was determined in the Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)], PDB code: 3qng:
Sodium binding site 1 out of 1 in 3qng
Go back to
Sodium binding site 1 out
of 1 in the Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)]
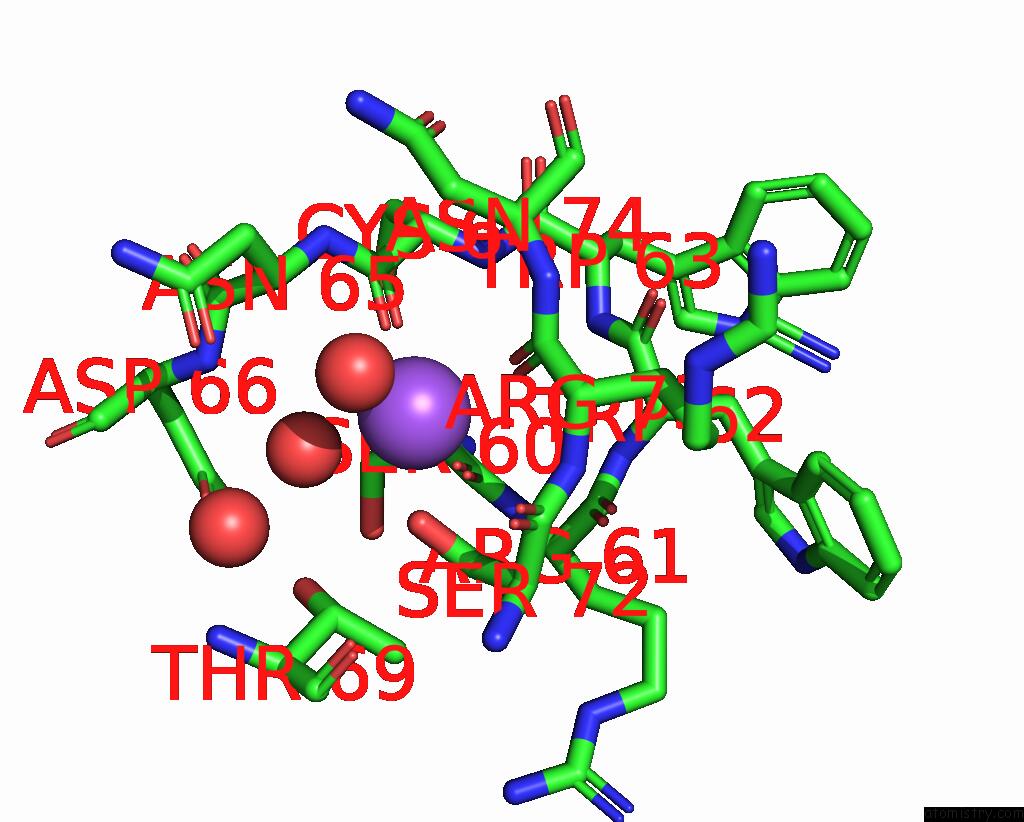
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of Crystal Structure Analysis of Lysozyme-Bound Fac-[Re(Co)3(L-Serine)] within 5.0Å range:
|
Reference:
F.Zobi,
B.Spingler.
Post-Protein-Binding Reactivity and Modifications of the Fac-[Re(Co)3]+ Core Inorg.Chem. V. 51 1210 2012.
ISSN: ISSN 0020-1669
PubMed: 22229733
DOI: 10.1021/IC2023314
Page generated: Mon Oct 7 12:35:33 2024
ISSN: ISSN 0020-1669
PubMed: 22229733
DOI: 10.1021/IC2023314
Last articles
Cl in 7YRACl in 7YR3
Cl in 7YQ7
Cl in 7YQ2
Cl in 7YQN
Cl in 7YR2
Cl in 7YQ9
Cl in 7YPZ
Cl in 7YNV
Cl in 7YNU