Sodium »
PDB 1z2u-1zud »
1zqs »
Sodium in PDB 1zqs: Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar)
Protein crystallography data
The structure of Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar), PDB code: 1zqs
was solved by
H.Pelletier,
M.R.Sawaya,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 20.00 / 3.30 |
| Space group | P 21 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 178.430, 57.723, 48.314, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 16.5 / n/a |
Sodium Binding Sites:
The binding sites of Sodium atom in the Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar)
(pdb code 1zqs). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total 2 binding sites of Sodium where determined in the Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar), PDB code: 1zqs:
Jump to Sodium binding site number: 1; 2;
In total 2 binding sites of Sodium where determined in the Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar), PDB code: 1zqs:
Jump to Sodium binding site number: 1; 2;
Sodium binding site 1 out of 2 in 1zqs
Go back to
Sodium binding site 1 out
of 2 in the Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar)
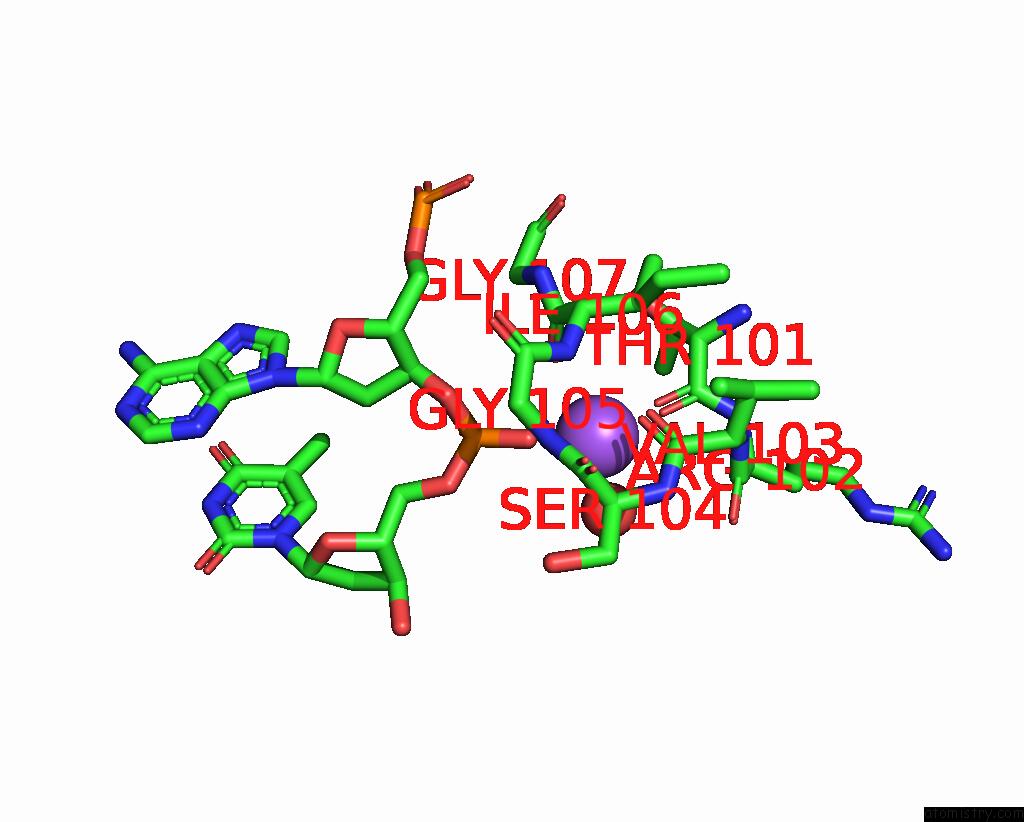
Mono view
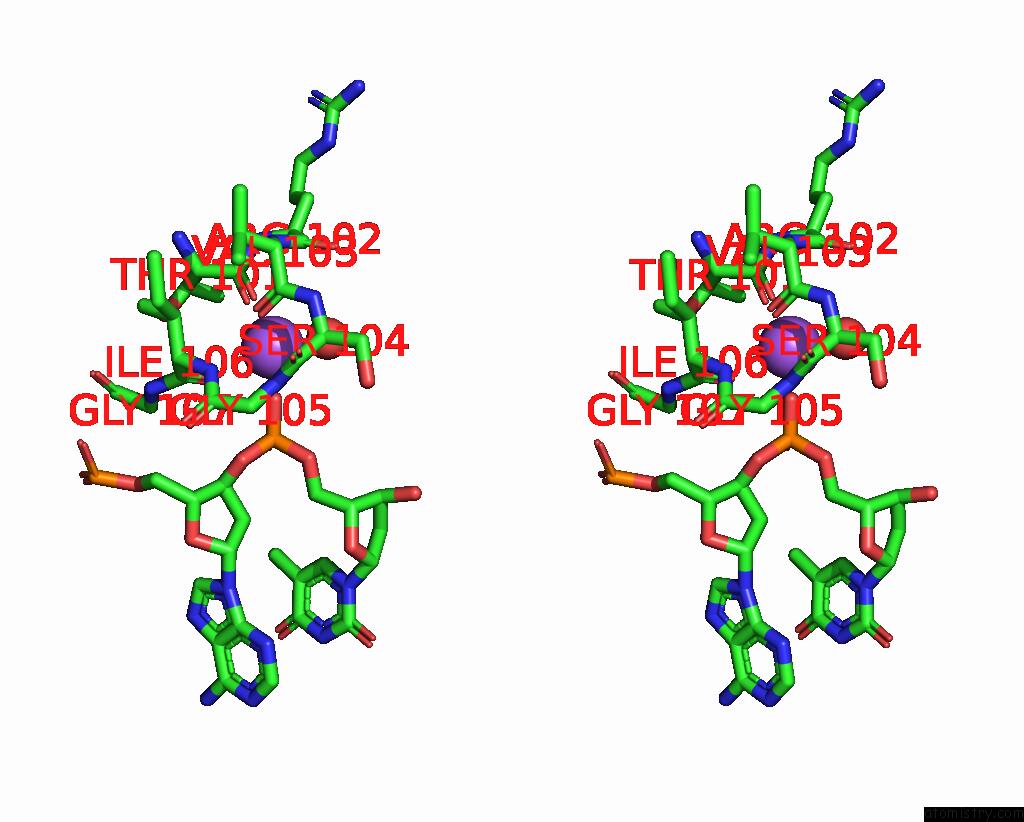
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar) within 5.0Å range:
|
Sodium binding site 2 out of 2 in 1zqs
Go back to
Sodium binding site 2 out
of 2 in the Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar)
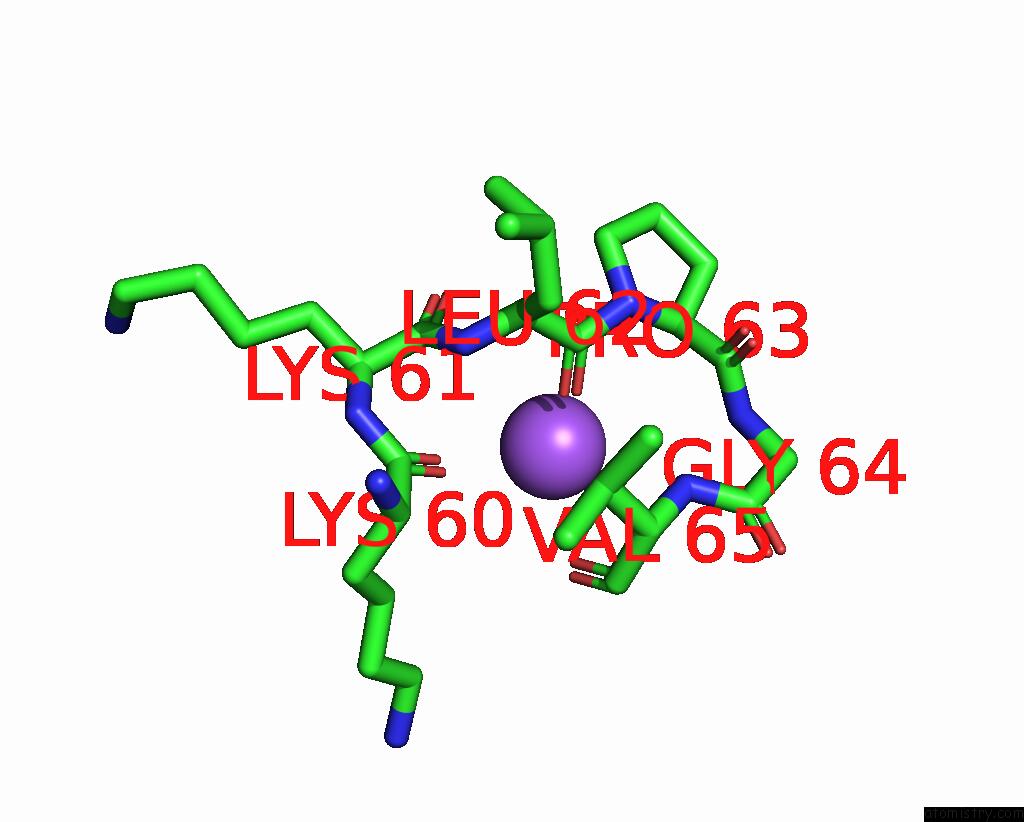
Mono view
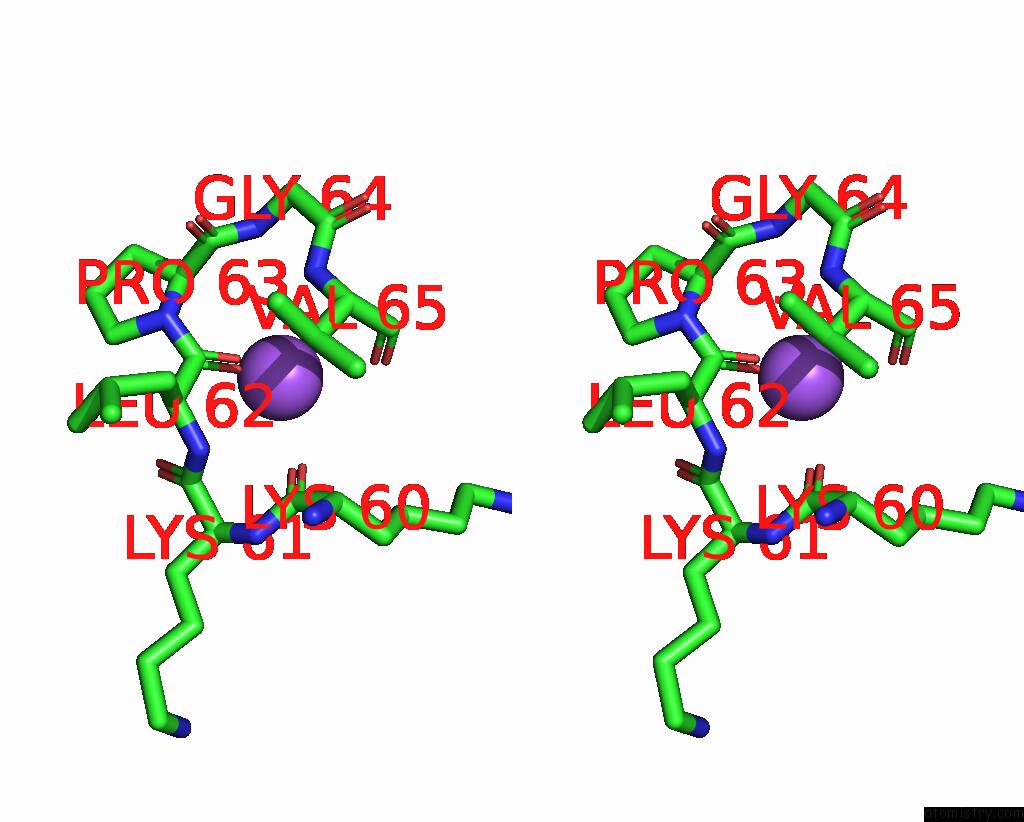
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 2 of Dna Polymerase Beta (Pol B) (E.C.2.7.7.7) Complexed with Seven Base Pairs of Dna; Soaked in the Presence of Tlcl (0.5 Millimolar) within 5.0Å range:
|
Reference:
H.Pelletier,
M.R.Sawaya.
Characterization of the Metal Ion Binding Helix-Hairpin-Helix Motifs in Human Dna Polymerase Beta By X-Ray Structural Analysis. Biochemistry V. 35 12778 1996.
ISSN: ISSN 0006-2960
PubMed: 8841120
DOI: 10.1021/BI960790I
Page generated: Sun Aug 17 10:00:42 2025
ISSN: ISSN 0006-2960
PubMed: 8841120
DOI: 10.1021/BI960790I
Last articles
Ni in 7Y2TNi in 7Y11
Ni in 7Y10
Ni in 7Y2S
Ni in 7Y2R
Ni in 7Y1S
Ni in 7WCY
Ni in 7XAR
Ni in 7XRG
Ni in 7XDP