Sodium »
PDB 1m90-1nji »
1m9m »
Sodium in PDB 1m9m: Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound
Enzymatic activity of Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound
All present enzymatic activity of Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound:
1.14.13.39;
1.14.13.39;
Protein crystallography data
The structure of Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound, PDB code: 1m9m
was solved by
R.J.Rosenfeld,
E.D.Garcin,
K.Panda,
G.Andersson,
A.Aberg,
A.V.Wallace,
D.J.Stuehr,
J.A.Tainer,
E.D.Getzoff,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 45.03 / 1.96 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 69.757, 90.782, 155.551, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.3 / 22.3 |
Other elements in 1m9m:
The structure of Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound also contains other interesting chemical elements:
| Iron | (Fe) | 2 atoms |
| Zinc | (Zn) | 1 atom |
Sodium Binding Sites:
The binding sites of Sodium atom in the Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound
(pdb code 1m9m). This binding sites where shown within
5.0 Angstroms radius around Sodium atom.
In total only one binding site of Sodium was determined in the Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound, PDB code: 1m9m:
In total only one binding site of Sodium was determined in the Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound, PDB code: 1m9m:
Sodium binding site 1 out of 1 in 1m9m
Go back to
Sodium binding site 1 out
of 1 in the Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound
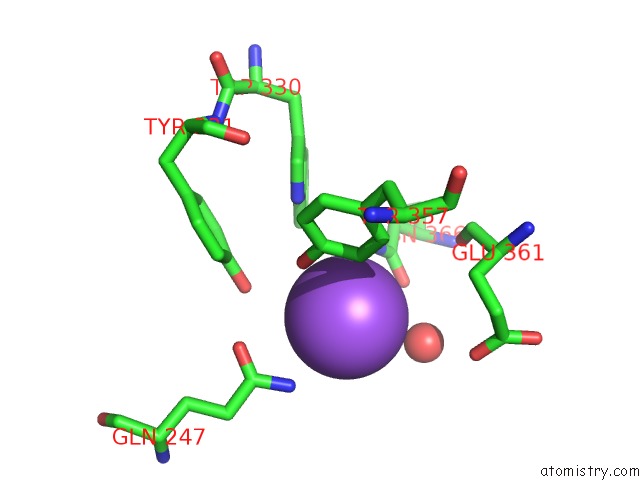
Mono view
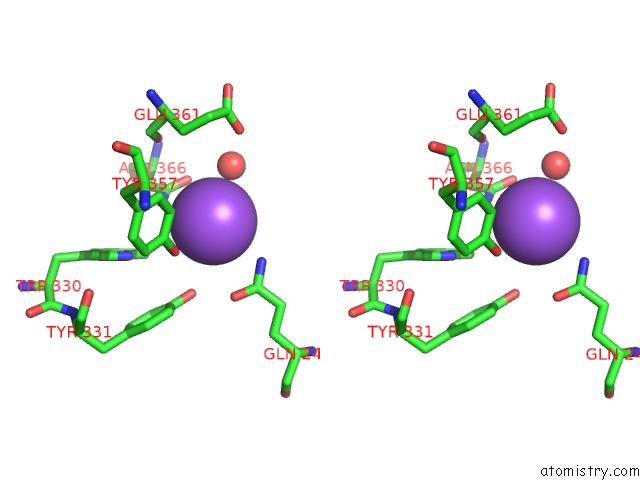
Stereo pair view

Mono view

Stereo pair view
A full contact list of Sodium with other atoms in the Na binding
site number 1 of Human Endothelial Nitric Oxide Synthase with 6- Nitroindazole Bound within 5.0Å range:
|
Reference:
R.J.Rosenfeld,
E.D.Garcin,
K.Panda,
G.Andersson,
A.Aberg,
A.V.Wallace,
G.M.Morris,
A.J.Olson,
D.J.Stuehr,
J.A.Tainer,
E.D.Getzoff.
Conformational Changes in Nitric Oxide Synthases Induced By Chlorzoxazone and Nitroindazoles: Crystallographic and Computational Analyses of Inhibitor Potency Biochemistry V. 41 13915 2002.
ISSN: ISSN 0006-2960
PubMed: 12437348
DOI: 10.1021/BI026313J
Page generated: Sun Oct 6 20:31:31 2024
ISSN: ISSN 0006-2960
PubMed: 12437348
DOI: 10.1021/BI026313J
Last articles
Fe in 2YXOFe in 2YRS
Fe in 2YXC
Fe in 2YNM
Fe in 2YVJ
Fe in 2YP1
Fe in 2YU2
Fe in 2YU1
Fe in 2YQB
Fe in 2YOO